When scientists first cracked the genetic code, they expected a simple story: DNA makes RNA, and that RNA, known as messenger RNA, makes proteins. Proteins would do all the important work — building tissues, fighting infections, digesting food.
But when the DNA of our genome was finally sequenced, researchers encountered a head-scratcher: The 20,000-plus genes that carry instructions for making our proteins account for less than 2 percent of our DNA. What was the rest of it good for?
For years, the remaining 98 percent was dismissed as “junk DNA” — evolutionary debris, filler. But as sequencing technology improved, a startling picture emerged. Our cells were busy making RNA copies of all those expanses, not just making messenger RNA — or mRNA — from the protein-coding genes. They were churning out vast quantities of RNA molecules with no known purpose.
The question became: Why would cells waste so much energy on copying that junk?
Today, however, the importance of this non-coding RNA — the catchall term for RNA molecules that don’t carry instructions for proteins — is undeniable. Non-coding RNAs turn out to regulate everything from embryonic development to immune responses to brain function. They help determine which genes get turned on and off, and when. They can promote cancer or suppress it.
“Every cell is probably a snowflake, and it’s due to the diversity of RNAs that is being produced in the genome — and that diversity does not come from mRNA,” says John Rinn, a regenerative biologist at the University of Colorado Boulder who has spent his career studying these molecules.
Here, Knowable Magazine profiles some of the major classes of non-coding RNAs that scientists have identified so far.
MicroRNAs: Tiny molecules, big effects
A key discovery in upending assumptions about non-coding RNA came from an itty-bitty source: a transparent worm called Caenorhabditis elegans. In 1993, developmental biologist Victor Ambros and geneticist Gary Ruvkun identified a peculiar gene in the worm that produced a tiny RNA molecule — just 22 nucleotides long (nucleotides are the building blocks of RNA and DNA). That RNA, it turned out, could keep other genes from making proteins. The scientists had found microRNA.
For years, the finding seemed like a curiosity specific to worms. Then Ruvkun found microRNAs in flies and fish, and other researchers identified them in plants and people. We now know that humans have more than 2,000 different microRNAs, and these molecules help control about a third of all protein-coding genes. For their discovery, Ambros and Ruvkun received the 2024 Nobel Prize in physiology or medicine.
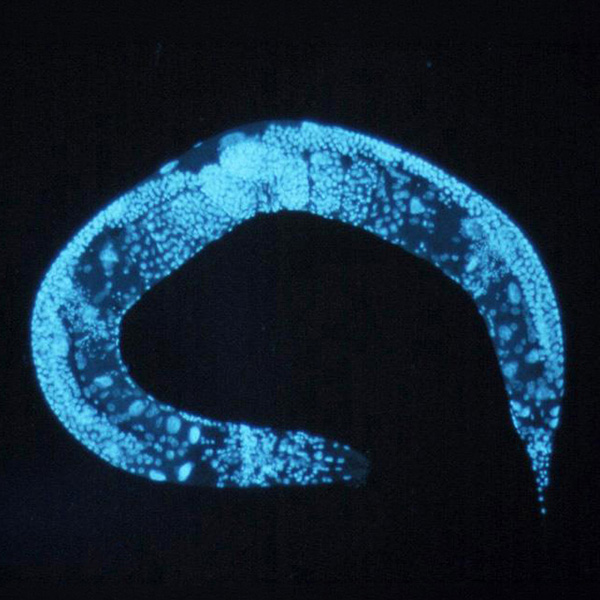
Non-coding RNA was once thought to be useless. A discovery that began to change that view came from a tiny worm called Caenorhabditis elegans (shown here in blue). In 1993, Victor Ambros and Gary Ruvkun identified a gene in the worm that produced a tiny RNA molecule — just 22 nucleotides long — that could silence other genes.
CREDIT: NIH / WIKIMEDIA COMMONS
MicroRNAs work by interacting not with the genes themselves but with the mRNA intermediaries that carry genetic instructions from DNA to the protein-making machinery. When a microRNA latches onto an mRNA, it either blocks the creation of a protein or triggers the mRNA’s destruction. A single microRNA can regulate hundreds of different genes, making these tiny molecules master controllers of cellular processes.
“What microRNAs do, is they add a level of precision over exactly how much of the various proteins are being made for individual cell types,” explains Kenneth S. Kosik, a neuroscientist at the University of California, Santa Barbara. That precision seems especially critical in the brain, which contains thousands of distinct cell types — more than any other organ. “In order to develop and to sustain this diversity of cells, having a corresponding diversity of microRNAs seems to be very important,” Kosik says.
MicroRNAs also circulate stably in the bloodstream, making them promising biomarkers for disease. Kosik’s team identified one called microRNA-21 that is overproduced in patients with a variety of cancers, including glioblastoma, an aggressive brain cancer. Pharmaceutical companies are also developing microRNA-based drugs, as well as molecules that block certain microRNAs, for conditions ranging from hepatitis to heart failure.
“We need to understand all their targets, how to control them,” Kosik says.
Long non-coding RNAs: The genome’s dark matter
If microRNAs are tiny, long non-coding RNAs are giants. Typically defined as any non-coding RNA 200 nucleotides or longer, they can stretch to tens of thousands of nucleotides in length. And there are lots of them in the human genome — 36,000 and probably many more, eclipsing the number of protein-coding genes.
The most famous long non-coding RNA — XIST — was discovered in 1991, and it performs a crucial role in females. Since they carry two X chromosomes, one of the pair must be silenced to avoid an imbalance of the genes that are carried on the X. XIST accomplishes this feat by coating the entire chromosome and then attracting proteins that inactivate most of the genes. “You can think of them as acting as scaffolds, where they can bring in other binding partners,” says Susan Carpenter, a cell and molecular biologist at the University of California, Santa Cruz.
And XIST is just one example. “Each long non-coding RNA can do different things,” Carpenter says. Some boost the activity of neighboring genes, others help to control when and where proteins are made, and still others carry instructions for making peptides — very short strings of amino acids — that can, for instance, regulate inflammation.
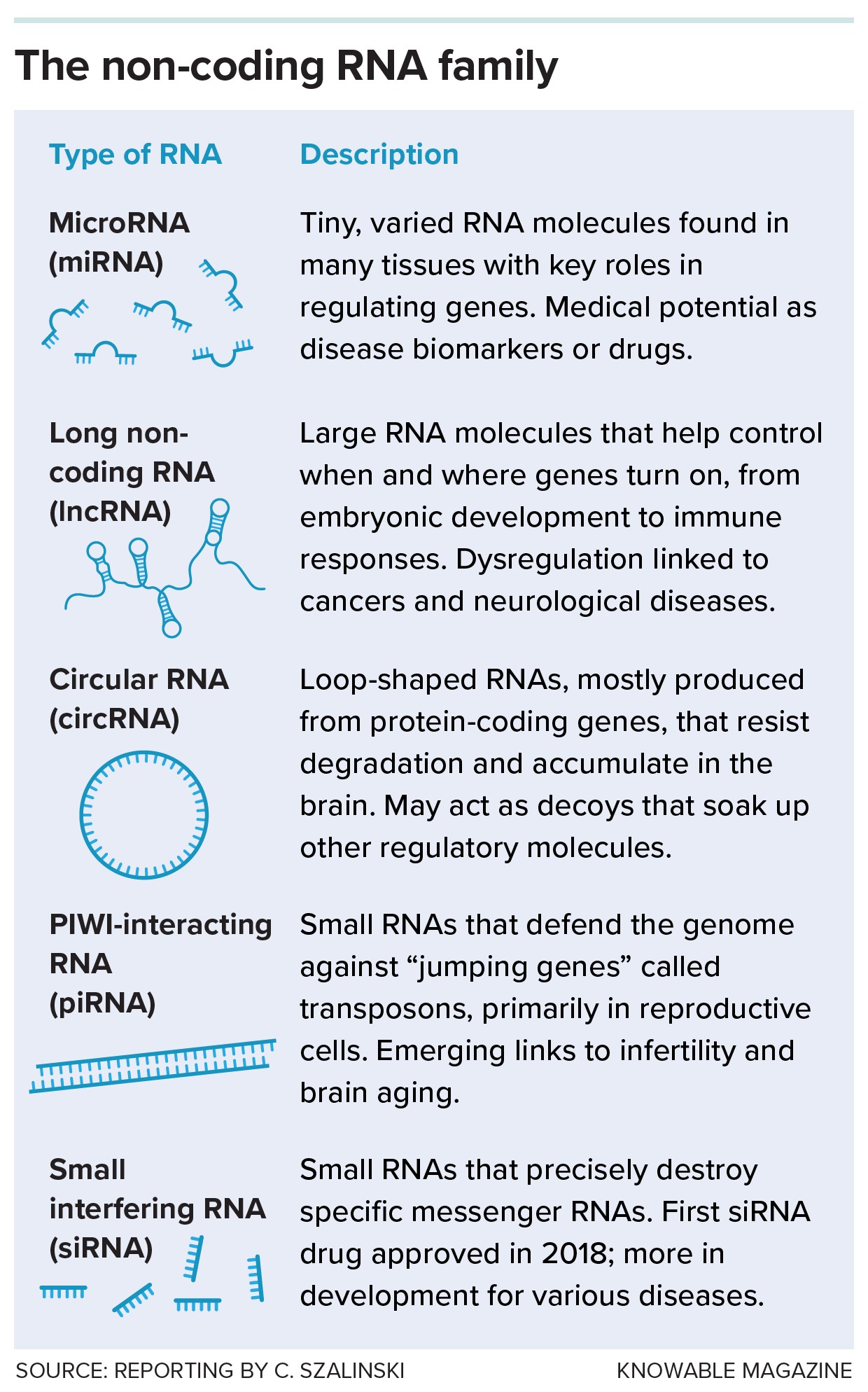
Five types of non-coding RNAs and their roles in the body. Unlike messenger RNA, these molecules don’t make proteins — instead, they regulate genes, defend the genome and perform other essential tasks.
Long non-coding RNAs are starting to matter clinically. In 2024, for example, researchers found that a long non-coding RNA called CHASERR is tied to a genetic condition: Having only one functional copy of the gene that makes CHASERR causes severe developmental disabilities. It appears, therefore, that some long non-coding RNAs are exquisitely sensitive to dosage.
Circular RNAs: Loops with a purpose
Traditional mRNA is linear — it has a beginning and an end, which the cell’s machinery needs in order to translate it into a protein. But some RNAs exist as circles, typically derived from the same genes that make linear mRNAs. When scientists first spotted them in the late 1970s, they assumed they were mistakes — splicing errors, cellular accidents.
Today, “there’s thousands and thousands of circles that have been identified,” says Jeremy Wilusz, an RNA biologist at Baylor College of Medicine. “They're particularly prominent in the brain, as well as some other tissues, but the brain is really where you see them the most.”
In fact, for hundreds of genes, the circular version is more abundant than the regular, linear mRNA copies that are made from a gene. And that is no accident. Because circular RNAs lack free ends, they’re resistant to the enzymes that normally chew up RNA molecules. This makes them exceptionally stable — they can persist in cells far longer than their linear counterparts, which are destroyed within hours.
That stability appears to serve a purpose. Circular RNAs can act as decoys, latching onto microRNAs or proteins and preventing them from doing their usual jobs, like regulating gene activity. One well-studied circular RNA called MALAT1 has been linked to several kinds of cancer — it’s abnormally abundant in tumors and seems to help them grow, though exactly how remains unclear. It appears to alter which genes are turned on or off, Wilusz says, but the details are still being worked out.
Emerging research links amounts of specific circular RNAs to fertility, brain aging and several cancers. Some companies are exploring whether circular RNAs could serve as more durable alternatives to the messenger RNA technology that is used in Covid-19 vaccines.
This video introduces the many types of non-coding RNAs, including some not covered in this article, and explains how they differ from the messenger RNAs that carry instructions for making proteins.
CREDIT: HENRIK’S LAB
piRNAs: The genome’s ancient immune system
In 2006, RNA molecular geneticist Nelson Lau and several other research groups independently discovered a new class of small RNAs that turned out to be the genome’s defense system against an ancient enemy.
The molecules, known as PIWI-interacting RNAs, or piRNAs, guard against entities called transposons (sometimes called “jumping genes”) that can copy themselves and insert those copies elsewhere in the genome. Transposons make up nearly half of all human DNA, and if left unchecked, their jumping could wreak havoc, disrupting essential genes and causing mutations.
The transposons are especially active inside the testes and ovaries, explains Lau, now at the Boston University Chobanian & Avedisian School of Medicine. Fortunately, he adds, “in our same germ cells, we also express a huge amount of these piRNAs.” When a transposon jumps into particular parts of the genome, the cell produces piRNAs from that new material; and these can thereafter recognize and silence that transposon and its relatives. In that sense, Lau says, piRNAs are “our very first genetic immune system to defend against viruses that want to propagate …through the germline.”
Lau’s lab has extended this work to mosquitoes, investigating whether this natural antiviral system could be harnessed to make mosquitoes resistant to the pathogens they transmit to humans. And piRNAs are also in the brain, where transposon activity increases with age. Lau is studying how failing transposon control might contribute to neurodegenerative diseases like Alzheimer’s and Parkinson’s.
siRNAs: From lab tool to medicine cabinet
Small interfering RNAs, or siRNAs, were discovered in plants and worms in the late 1990s. Like microRNAs, they’re small — typically 21 to 23 nucleotides — and they work by binding to particular messenger RNAs and targeting them for destruction. By so doing, they too help control transposons and defend against viruses.
Their high specificity made them immediately attractive for therapeutics. Scientists realized they could design synthetic siRNAs to silence virtually any gene — a powerful tool for treating diseases that were caused by harmful, malfunctioning proteins. In 2018, the first siRNA drug, Patisiran, was approved for a rare hereditary disease called transthyretin amyloidosis.
The technology faces challenges, among them how to deliver siRNAs to the right tissues in the body and how to avoid immune reactions. But new chemical modifications and drug delivery systems are showing promise, and the pipeline of siRNA drugs continues to grow.
A new view of the genome
As researchers learn more, the picture of non-coding RNAs that’s emerging is one of dizzying complexity. MicroRNAs regulate long non-coding RNAs. Long non-coding RNAs can sponge up microRNAs. Circular RNAs arise from the same genes that produce messenger RNAs. piRNAs silence transposons that have shaped our genome for millions of years.
And all of this regulation happens on top of the protein-coding genes and mRNA that were once thought to be the whole story.
Plus, scientists have studied only a fraction of the non-coding RNAs known to exist — perhaps just 3 percent, Carpenter says — leaving a great deal still to discover.